Autophagy is the controlled digestion and degradation of damaged and old cell and its components. Form the Greek origin, “auto” mean “self” and “phagein” refers to “eat”. In the process, the cell can digest its own components by enclosing them in a membrane forming sac-like vesicles. These are later transported to lysosome. Lysosome is the recycling compartment of the cell and digests the old and damaged components of cell.
Yoshinori ohsumi was awarded the Nobel Prize in the year 2016 for his work on autophagy in the field of physiology & medicine.
Work of Yoshinori ohsumi
Professor Yoshinori identified the mechanism and the genes essential for the phenomenon of autophagy in the yeast cell that helped to understand this phenomenon in human body cells. His work explained the phenomena and mechanism of cellular component’s recycling.
Christian De Duve discovered a specialized cellular organelle of the cell that contains enzyme for the celluler protein’s digestion in mid 1950’s that was termed as “lysosomes”. He used the term “autophagy” later after observing and analyzing some cellular process. The cells use these processes for the digestion of cellular proteins and even the cell components as well. During the process, specified vesicles were observed in the cell that was defined as “autophagosomes.”
Yoshinori conducted a series of experiments for detailed analysis. He reasoned:
“If the degradation process of the cell could be disrupted in the vacuole during the process of active autophagy in cell; then there will be autophagosomes accumulation within the cell vacuole. Moreover, it will then be visible under electron microscope.”
Researcher cultured and mutated yeast cells in lab. The mutated lacked the derivative enzymes of vacuole. Moreover, the cellular starvation stimulated the process of autophagy.
Within the few hours, the vacuoles filled with small undigested vesicles. This experiment proved that the phenomenon of authophagy existed in the yeast cells. Also these experiments identified the key regulatory genes of process.
Regulatory genes and proteins
Professor Ohsumi identified the regulatory genes for autophagy. Researchers identified 15 regulatory genes after analyzing and studying thousands of yeast mutants. Later, the proteins encoded by these genes were identified also.
The whole series of experiments suggested that some specific proteins control cellular autophagy. Each of them regulates and controls some specific step for the formation of autophagosomes and eventually the whole process.
Professor Yoshinori also studied the function of the proteins encoded by key autophagy genes. For example, the Becilin1 protein. He analyzed how the stress signals were responsible for initiating the mechanism by which proteins and protein complexes regulate each stage of autophagosome formation.
The protein complexes are:
- Comples 1 that consists of the class of proteins III PI3K/Vps34, Atg6/Beclin1, Atg14 and Vps15/p150.73.
- Complex 2 that consists of srine/threonine kinase Atg1/ULK1 that is a positive regulator of the whole process of autopaghy
Mostly, phosphorylation of certain proteins by protein kinases activates the proteins or the enzyme for certain steps and also the dephosphorylation may deactivate that protein.
Basic mechanism of Autophagy
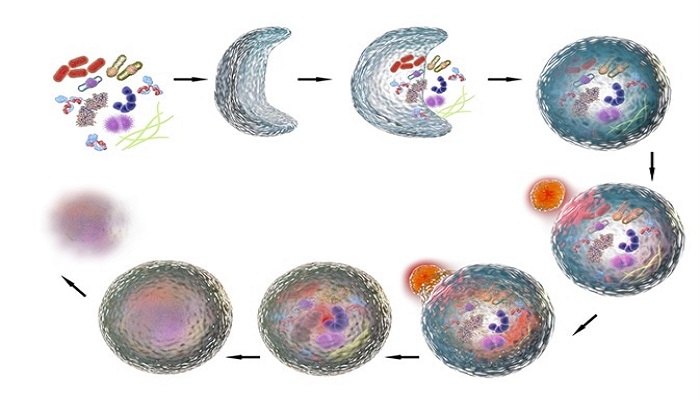
The complete process consists of four distinct phases:
- Induction and phagophore formation
- Autophagosome formation
- Fusion with lysosome
- Degradation by lysosomal enzymes
Phagophore describes a double membranous sac-like structure which forms at the initial stages of autophagy. The phagophore elongates similar to ubiquitin pathway and forms an autophagosome that attaches or fuses with the lysosome in the next step. Later, lysosomal enzymes derive a series of acid hydrolyses reactions, for degradation of the protein in autophagosome.
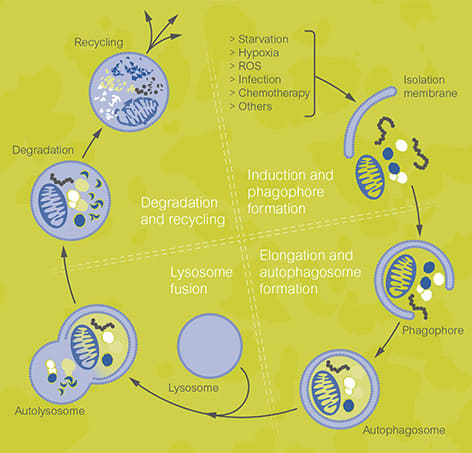
“Basic mechanism and steps involved in autophagy”
The whole process relies upon the autophagy-related genes and the protein complexes encoded by these genes. Certain “permeases” are also participate in this process. Permeases are transport or carrier proteins for the autophagosome’s transport to the lysosome.
Significance of Autophagy
- The process of autophagy is a source of feul and building blocks for renewal of cell components. So in case of starvation and stress, this process is essential.
- Similarly in case of infection, it also helps to eliminate the bacterial components and the inflammation causing substances. Autophagic degradation of the infectious particles is termed as “xenophagy.”
- Also, the process is vital during the developmental stages of embryo and cell differentiation.
- Cells use this phenomenon to remove and eliminate the cellular debries and the damaged proteins.
Association with Diseases
1. Parkinson’s disease
Mutations in autophagy genes may result in genetic disorders and cancer as well. Parkinson’s disease is a neurodegenerative disorder. This is characterized by the decline in brain functions and memory loss. Autophagy-related genes mutation may be one of the causative agents of Parkinson disease because of the accumulation of certain cellular wastes and their incomplete degradation.
2. Lysosomal storage disease
The presence of undigested macromolecules in cell due to disrupted autophagy process results in lysosomal storage disease.
3. Cancer
In case of cancer, the process of autophagy prevents the tumor formation in cell. The abnormal tumor cells use autophagosome for their own protection when develop. So initially, autophagy acts as tumor suppressor and then the cancer cells use it for their survival.
Researches are now focusing on the aspect that the self-induced and targeted autophagy can serve as treatment strategy for cancer.
Bottom line
On the basis of the given facts, we can summarize the significance of autophagy for the normal cellar functioning as it provides the basis for degradation of the cellular waste components and their renewal. Also the bacterial infections are prevented as the infectious particles are digested and degraded by xenophagy.